Subjects
Most of the procedures are the same as in ref. 26, but will be summarized below.
Two rhesus monkeys (Macaca mulatta; 1 female; 5–9 kg), M1 and M2, were used for the high-resolution fMRI study. They had previously participated in several other studies26,27,56,57. Another male rhesus monkey, M3 (6.5 kg), was used for the electrophysiological experiments. The latter subject also participated in several previous studies15,24. Animal care and experimental procedures were performed in accordance with the National Institute of Health’s Guide for the Care and Use of Laboratory Animal, the European legislation (Directive 2010/63/EU) and were approved by the Animal Ethics Committee of the KU Leuven. Weatherall reports were used as reference for animal housing and handling. All animals were group-housed in cages sized 16–32 m3, which encourages social interactions and locomotor behavior. The environment was enriched by foraging devices and toys. The animals were fed daily with standard primate chow supplemented with fruits, vegetables, bread, peanuts, cashew nuts, raisins, and dry apricots. The animals were exposed to natural light and additional artificial light for 12 h every day. On training and experimental days, the animals were allowed unlimited access to fluid through their performance during the experiments. Using operant conditioning techniques with positive reinforcers, the animals received fluid rewards for every correctly performed trial. During non-working days, they received water in their living quarters. Throughout the study, the animals’ psychological and veterinary welfare was monitored daily by the veterinarians, the animal facility staff and the lab’s scientists, all specialized in working with non-human primates. The animals were healthy at the conclusion of our study. M1 and M2 are currently still employed in other studies.
To improve the sensitivity of sub-millimeter resolution (f)MRI, 8 (M1), or 10 (M2) channel phased-array receive coils were embedded in an MRI-compatible headpost above the skull23. M3 was also implanted with a magnetic resonance (MR) compatible headpost but lacked an embedded phased-array receive coil. M3 was equipped with a recording chamber targeting the middle portion of the superior temporal sulcus (STS).
The monkeys were trained to maintain fixation within a 2 \(\times \,\)2° virtual window in the center of the screen while sitting in a sphinx position inside a plastic primate chair, and with their heads constrained by a plastic headpost. Fluid rewards were contingent upon fixation behavior and keeping both hands positioned on response keys within the response box in front of the chair -which was monitored using infrared beams. Before each fMRI scan, 8–11 mg/kg monocrystalline iron oxide nanoparticle (Molday ION, BioPAL) was injected via the femoral/saphenous vein to improve the contrast-to-noise ratio and to reduce the contribution of superficial draining veins25. To mitigate the risk of iron accumulation, 1 g/day deferoxamine mesylate (Desferal, Novartis; intramuscular injection) and 60 mg/kg/day deferiprone (Ferriprox, ApoPharama; oral administration) were administered immediately after the scan, and we continued this iron chelation for 4–20 days, until serum iron and ferritin level returned approximately to normal ranges.
(f)MRI acquisition
High resolution fMRI (M1 and M2)
A 3 T Siemens PrismaFit scanner was used at the KU Leuven. For sub-millimeter (0.6 mm isotropic voxel size) functional measurements, we used a 2D simultaneous multiple slice gradient echo planar imaging sequence using the 10- or 8-channel implanted receive coils and a custom-built local single loop transmit coil to cover the whole brain [echo time (TE) = 22/21 ms; accelerated multiband (MB) = 2; acceleration factor = 3; in-plane field of view (FOV) 84 \(\times\) 84 mm; matrix size 140 \(\times\) 140; slices = 74 (M1)/80 (M2), adjusted according to individual brain size; flip angle (\(\alpha\)) = 90°]. Repetition time (TR) = 3000 ms in all experiments except for resting state experiments (TR = 2850 ms).
During a separate session under ketamine-medetomidine anesthesia, we obtained template EPI images using the same sequences, coils, and positions of the subjects, as during the awake behaving fMRI sessions. The corresponding field maps, acquired within the same session, were used to correct for EPI distortions caused by magnetic field inhomogeneity [0.6 \(\times\) 0.6 \(\times\) 0.7 mm voxel size; TR = 917 ms; \({{\rm{T}}}{{{\rm{E}}}}_{1}\) = 6.48 ms, \({{\rm{T}}}{{{\rm{E}}}}_{2}\) = 8.94 ms; in-plane FOV 84 \(\times\) 84 mm; matrix size 140 \(\times\) 140; slices = 66 (M1)/70 (M2), adjusted according to individual brain size; \(\alpha\) = 55°]. To achieve a better registration between the EPI template image and the anatomical reference images, we also acquired T1-w 3D magnetization prepared rapid gradient echo (MPRAGE) images (0.6 mm isotropic voxel size; TR = 2700 ms; TE = 3.8 ms; in-plane FOV 154 \(\times\) 125 mm; matrix size 256 \(\times\) 208; slices = 144; \(\alpha\) = 9°; inversion time (TI) = 850 ms) in the same session during which the animals were anesthetized. We used these as intermediate images for registration of images acquired in the awake fMRI sessions and the high-resolution T1-weighted images (acquired in a different session using anesthesia).
To visualize the functional results, we acquired high-resolution (0.4 mm isotropic voxel size) T1-w and T2-w images when the subjects were under ketamine-xylazine (M1) or ketamine-medetomidine anesthesia (M2), using a custom-built local single-loop receive coil and the body transmit coil of the scanner. 13 and 11 T1-w 3D images were acquired respectively for subjects M1 and M2 using a 3D MPRAGE sequence [TR = 2700 ms; TE = 3.5 ms; in-plane FOV 104 \(\times\) 128 mm; matrix size 250 \(\times\) 320; slices = 208; \(\alpha\) = 9°; TI = 882 ms]. 5 and 4 T2-w 3D images were acquired from M1 and M2 with the same in-plane FOV and matrix size as the T1-w images, to reduce field inhomogeneities in the anatomical images and to extract dura and blood vessels from pial surfaces58. The T2-w images were acquired using a sampling perfection with variable flip angle turbo spin-echo (SPACE) sequence (TR = 3200 ms; TE = 456 ms; total turbo factor = 131; echo spacing = 6 ms; no fat suppression).
Low-resolution fMRI for the electrophysiology monkey M3
Details of the fMRI procedure, data analysis and results are provided in refs. 15,24 and are very similar to those described above for the high-resolution fMRI experiments of M1 and M2 -except that a 3T Siemens Trio scanner was used and the spatial resolution was 9 times lower (1.95 mm3 voxels) than in the high-resolution experiments (0.22 mm3 voxels). Specifically, we used a gradient-echo single-shot echo planar imaging (EPI) sequence (TR = 2000 ms; TE = 17 ms; \(\alpha\) = 75°; matrix size 80 \(\times\) 80; slices = 40; 1.25 mm isotropic voxel size). The functional images were co-registered with a high-resolution (0.4 mm isotropic) T1-w 3D anatomical image of the monkey’s individual brain (see above), serving as a template.
Experimental design and stimuli
The stimuli in the scanner (M1-3) were projected using a Barco LCD projector at 60 Hz refresh rate and 1400\(\,\times\) 1050 resolution onto a translucent screen located at 57 cm from the subjects’ eyes. An eye-tracking system based on infrared corneal reflection (ISCAN, 120 Hz) was used to monitor eye movements. Only those runs in which the subjects fixated within a 2° \(\times\) 2° fixation window centered on the middle of the screen for more than 90% of a run were retained for further analysis. Moreover, high-resolution fMRI monkeys M1 and M2 were required to keep both hands on two response keys within a response box in front of the chair. This helped to significantly reduce motion-induced susceptibility artifacts.
fMRI design
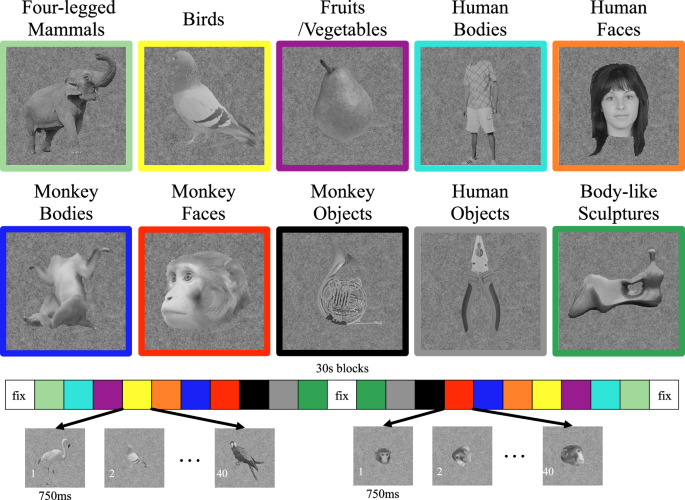
A block design was conducted (as in ref. 26). Each block lasted 30 s for M1 and M2, and 20 s for M3 (see Fig. 1). There were multiple run orders in each experiment, counterbalanced across runs such that each condition occurred equally often in each serial position. Each run started and ended with a blank fixation block, in which a uniform grey background was presented with the same luminance as the mean of the images shown in the other conditions.
Category localizer
The same stimuli were used in the high-resolution fMRI experiments in M1 and M2, and the low-resolution fMRI experiments in M3. There were ten classes of achromatic images—monkey and human bodies (excluding the head), monkey and human faces, four-legged mammals, birds, manmade objects (matched either to the monkey or to the human bodies), fruits/vegetables, and body-like sculptures (by the British artist H. Moore). Each class consisted of 20 images, which were previously used in the fMRI study of 15,24. Examples of the stimuli are shown in Fig. 1, while the full stimulus set together with details about the stimuli can be found in ref. 24.
We made every effort to equate the low-level image characteristics, such as mean luminance, mean contrast, and aspect ratio, across the different stimulus classes. The mean aspect ratio of the monkey and human bodies differed since the upright human bodies tend to be more elongated than the monkey bodies. This was controlled for by using two classes of manmade objects—one matching the aspect ratio of the monkey bodies (monkey objects) and another one matching the aspect ratio of the human bodies (human objects). The images were resized so that the average area per class was matched across all classes, except for the human objects and human bodies, but still allowing some variation in area (range: 3.7° to 6.7° (square root of the area)) within each class. The mean vertical and horizontal extent of the images was 8.3° and 6.7° of visual angle, respectively. The images were embedded into pink noise backgrounds having the same mean luminance as the images and which filled the entire display (height \(\times\) width: 30° \(\times\) 40° of visual angle). Each image was presented on top of 9 different backgrounds that varied randomly across stimulus presentations. The stimuli were gamma corrected.
Each category contained 20 images from multiple individuals, each image was shown twice and lasted for 750 ms (M1 and M2) or 500 ms (M3) in every block. Each run lasted for 705 s (M1 and M2) or 430 s (M3), containing 235 (M1 and M2) or 215 (M3) functional volumes. 30 runs were acquired for M1, 55 runs for M2, and 28 runs for M3.
Resting-state fMRI
The subjects were asked to maintain fixation (within a 2 \(\times\) 2° window) at a small red fixation point on a uniform gray background during the entire run. Each run lasted 658 s, containing 231 functional volumes. 28 runs and 37 runs were acquired for M1 and M2, respectively.
Electrophysiological recordings
Standard single-unit recordings were performed with epoxylite-insulated tungsten microelectrodes (FHC; in situ measured impedance between 1.3 and 1.6 MΩ) using techniques as described previously59. Briefly, the electrode was lowered with a Narishige microdrive into the brain using a stainless steel or an MR-compatible (when a position verification scan was performed after recording) guide tube that was fixed in a standard Crist grid positioned within the recording chamber. After amplification and filtering between 540 Hz and 6 KHz, spikes of a single-unit were isolated online using a custom amplitude- and time-based discriminator.
The recording grid locations were defined so that the electrode targeted the left MSB body area in M3. Before the recordings started, we performed a structural MRI and visualized long glass capillaries filled with the MRI-opaque copper sulfate (CuSO4) that were inserted into the recording chamber grid (until the dura) at predetermined positions. Then, the functional images (the contrast between the monkey bodies and monkey objects) of each monkey were co-registered with its anatomical MRI using the co-registration toolbox of SPM8 (Wellcome Department of Cognitive Neurology, London, UK) and the registration was verified by visual examination. Primary grid positions were selected for MSB recordings if the electrode would end in a voxel that was activated significantly more by monkey bodies than monkey objects and was not activated by monkey faces compared to monkey objects. Only neurons that were body-selective within MSB of M3 were included in the analysis, as this was also the (stringent) criterion for selecting the voxels in the high-resolution fMRI experiment in M1 and M2 (focusing on MSB) -see above. During the recordings, we verified the recording locations with 10 additional anatomical MRI scans. Four of these scans were performed immediately after recording sessions that targeted the body area, using an MR-compatible (fused silica; Plastics One, Roanoke, VA, US) guide tube with the electrode left in the cortex during the MRI scan. In all other scans we visualized long glass capillaries filled with copper sulfate that were inserted into the grid at recorded grid positions. The recording locations along the medio-lateral and antero-posterior dimensions were extrapolated from the trajectories of the imaged capillaries. The validity of the latter method to verify recording locations is supported by 4 MRI scans in M3 in which the electrode was imaged directly and was indeed shown to be present at the predicted location in the anterior-posterior and medial-lateral dimensions. The ventral-dorsal location of the electrode tip was verified in each recording session using the transitions of white and gray matter and the silence marking the sulcus between the banks of the STS.
Since the entire stimulus set (200 images) was too large to be presented in single cell recording sessions, neurons were searched while presenting half of the images (100 images) from the main stimulus set in a pseudo-random order. Stimuli were presented for 200 ms each with an inter-stimulus interval (ISI) of approximately 400 ms during passive fixation (fixation window size 2° \(\times\) 2°). The pink noise background was present throughout the task but refreshed together with the stimulus onset. Fixation was required in a period from 100 ms pre-stimulus to 200 ms post-stimulus. A trial was aborted when the monkey interrupted fixation in this interval. In the pseudo-randomization procedure, all 100 stimuli were presented randomly interleaved in blocks of 100 unaborted trials. Aborted stimulus presentations were repeated within the same block in a subsequent randomly chosen trial. ISIs within and between successive blocks were the same. Juice rewards were given with decreasing intervals (2000 ms to 1350 ms) as long as the monkeys maintained its fixation. All neurons were tested using this procedure and testing was continued when a response was notable in the on-line Peri-stimulus Time Histograms for at least one of the stimuli.
Retinotopic mapping experiment
The same stimuli as in refs. 27,57,60 were used for retinotopic mapping. Eccentricity and polar angle were mapped using phase-encoded annuli and wedges, which contained both dynamic monkey faces and walking humans. To mitigate phase errors caused by hemodynamic response delays, both expanding and contracting annuli, as well as clockwise and counter-clockwise rotating wedges, were used. The stimuli covered the central 0.25° to 12.25° of the visual field (radius). Each run included 4 stimulus cycles, with each cycle lasting 96 s. During a run, only one type of stimulus (e.g., a wedge or annulus moving in a specific direction) was displayed. A central fixation point was shown throughout, and the subjects were required to maintain fixation on this point. Only runs in which the subjects maintained fixation within a small virtual fixation window (2° \(\times\) 2° around the fixation point) for more than 90% of the time, while keeping their hands in a rest position, were used for further analysis. The data were analyzed exactly as described in ref. 27.
Data analysis
Stringent definition of face and body areas for the high-resolution fMRI experiment (M1 and M2)
Face and body areas were defined in each monkey’s native space. Face-selective area ML61 was defined using the conjunction of the following 3 contrasts (each with p < 0.05): monkey faces versus monkey objects, monkey faces versus fruits, and monkey faces versus monkey bodies. Body-selective area MSB24,62,63 was defined in a similar way using 3 different contrasts (each with p < 0.05): monkey bodies versus monkey objects, monkey bodies versus fruits, and monkey bodies versus monkey faces. As we aimed to be conservative in our definition of ML and MSB, we included only those voxels in these ROIs showing highly reproducible face or body selectivity. Therefore, we inclusively masked the voxels showing significant face or body selectivity across odd and even days for the 3 above-mentioned contrasts.
Mesoscale functional units (MFUs) in face and body areas and determining the optimal number of clusters
To estimate fMRI tuning in each individual voxel from the face and body areas, the effect size of each category versus fixation in units of percent signal changes were normalized across the 10 conditions by converting them to z-scores. A dissimilarity matrix was calculated based on pairwise Euclidean distances among all voxels within a face or body selective area. Then we performed hierarchical cluster analysis with Ward’s method on the dissimilarity matrix. The number of clusters was estimated based on the maximum differences in the trace of the within clusters pooled covariance matrix (Trace Cov W)64.
The Trace Cov W index quantifies the total within-cluster variance across all clusters, with a lower value indicating more compact clusters. By definition, this index decreases as the number of clusters increases because data points are divided into more groups, resulting in tighter and more compact clusters. The goal of this metric is to determine the optimal number of clusters by identifying the point where further increases in cluster no longer lead to a significant reduction in within-cluster variance, or where the decrease in variance between consecutive counts is most pronounced.
To further validate the number of clusters, we performed a permutation analysis of the Trace Cov W index and a consensus clustering analysis. For permutation testing, we randomly shuffled the data 10,000 times and calculated Trace Cov W indices for each permutation. We then compared them to the observed Trace Cov W index from the original data, to determine at which cluster count the drop in the Trace Cov W index (1st difference) is significantly larger than drops calculated from randomly permuted data. For consensus clustering, we performed hierarchical clustering on 1000 resampled subsets of the data (75% of the data each time). We then assessed the cluster stability and robustness for different values of K by calculating the proportion of times each pair of data points co-clustered across these iterations. From the resulting consensus matrices (values ranging from 0— indicating two data points were never clustered together, to 1—indicating they were always clustered together), we then evaluated the increase in clustering stability with increasing K by examining the change in area under the CDF of the consensus values.
FC analysis based on resting-state fMRI data
We used identical procedures as described in ref. 26. We included slice timing correction and the signal was high-pass filtered at 0.0025 Hz and low-pass filtered at 0.05 Hz. The signal from white matter and ventricle ROIs, the motion correction regressors, reward, and eye movement related regressors, and their first derivatives were used to regress out nuisance effects. No global mean regression was performed to avoid discarding any underlying neural components65. The representative time course of each MFU was obtained by averaging the signals across voxels. Pearson correlations between all possible pairs of MFUs were calculated to measure the functional connectivity. The correlations were converted to z-scores by Fisher’s r-to-z transformation per run66 to improve normality.
Quantitative tests comparing interhemispheric functional connectivity
Considering that resting-state signal fluctuations in cortical regions tend to be positively correlated between homologous regions from left and right hemispheres67, even in the absence of direct anatomical connections68, we predicted that the same holds true for MFUs of the same type across the two hemispheres. Therefore, we tested whether FC between MFUs belonging to the same cluster ( = same-type MFUs) of a large and well-established face area (ML), yet across hemispheres, is stronger than between MFUs (across hemispheres) belonging to different clusters of the same face area ( = different-type MFUs). To generalize this result, we also performed the same analyses for MFUs of the body area MSB. To test inter-MFU functional connectivity quantitatively, we first calculated the FC strengths from the same- and different-type MFUs across hemispheres for each resting-state run. We then performed a pairwise t-test to compare FC strengths between same- and different-type MFUs across all the resting state runs. Note that (i) two MFUs can be different in size, (ii) smaller units tend to exhibit lower SNR, and (iii) FC strengths can be biased depending on the size of these functional units. To control for this bias, we performed a permutation test (10,000 times of permutations) and calculated the probability that the median value of FC differences between same- or different-type MFUs was higher than the median value when the same voxels were randomly assigned to different MFUs.
Comparison between MFUs based on fMRI and functional clusters based on single-cell recordings in body area MSB
To test whether similar sub-areal functional clusters appear at single-cell level, we re-analyzed single-unit data15, which were recorded in fMRI-defined MSB of another animal (M3). We only included the single-unit data in our analysis, which showed body selectivity for the conjunction of the same contrasts as used in the high-resolution fMRI study (monkey bodies versus monkey objects, monkey bodies versus fruits, and monkey bodies versus monkey faces, each with p < 0.05). Each cell’s average (net) spike response to the 100 stimuli was normalized by converting them to z-scores, and a dissimilarity matrix was calculated based on pairwise Euclidean distances across all cells. Then we performed hierarchical cluster analysis with Ward’s method on the dissimilarity matrix. As the number of data points (i.e., 98 cells) is smaller than the number of features (i.e., 100 stimulus images), Trace Cov W index cannot be used to determine the number of clusters. We therefore chose the number of clusters based on the Trace Cov W index calculated from averaged single-cell responses at category level. Finally, nonparametric Spearman correlation coefficients were calculated on the average response profile between each fMRI and single-cell clusters. Since single-cell and fMRI data were recorded from different subjects, we pooled data from the 2 fMRI subjects with the goal to reduce individual biases. We first calculated average profiles for each cluster from each individual and then calculated the mean of the corresponding profiles from the two individuals. Nonparametric Spearman correlations were used to calculate the distance matrix, and a hierarchical cluster analysis with the complete linkage method was conducted to objectively match corresponding clusters obtained by either fMRI or single-unit recordings.
Evaluating the impact of eccentricity biases or face/body-selectivity on MFU clustering
To determine whether MFUs could be distinguished based on eccentricity differences, we computed correlations between Euclidean distance matrices derived from eccentricity maps and those from full object response profiles. This analysis quantified how much of the variance among the 3 clusters could be attributed to eccentricity. Eccentricity was estimated using a general linear model applied to retinotopic mapping data. Phase values for the annuli served as proxies, with the fovea assigned a phase of zero and the most peripheral eccentricity a phase of 2π. These phase values were z-normalized across voxels before computing Euclidean distances. Only voxels exhibiting significant eccentricity biases (p < 10−3) were included in the analysis. Object response profiles were derived as described previously for hierarchical analysis, with effect sizes calculated as percent signal change of each category relative to fixation. These values were z-normalized across conditions before computing Euclidean distance matrices. To assess the reliability of these distance measurements, we performed test-retest analyses. Data acquired on different days from each experiment were split evenly into two independent datasets, and the consistency of distance measurements from both eccentricity and object response profiles was evaluated using test-retest correlation analysis and the Spearman-Brown formula.
We conducted a similar analysis to quantify the contribution of face selectivity (for ML) and body selectivity (for MSB) to the variance among the 3 clusters. Face selectivity was estimated using t-values from an “all faces versus all bodies” contrast, while body selectivity was calculated from the reverse contrast. These t-values were z-normalized across voxels before computing Euclidean distances. To evaluate the variance explained by face and body selectivity, we computed correlations between Euclidean distances derived from these selectivity measures and those from the full object response profiles. Additionally, we computed split-half analysis, following the same procedure used for eccentricity, to assess the reliability of these distance measurements.
Quantify the sharpness of MFU boundaries
To quantify whether object response profiles change gradually or sharply across MFU borders, we compared Euclidean distances between response profiles of neighboring voxels. For each voxel located at an MFU border, we calculated the Euclidean distance between its response profile and those of its neighboring voxels within the same MFU (“within” condition), as well as those of its neighboring voxels belonging to a different MFU (“across” condition). The two sets of distances were compared using a Wilcoxon rank-sum test.
Quantify the response similarity spatial fall-off pattern
Since MFUs were clustered based on Euclidean distances, we used 1- Euclidean distance as an index of response similarity between voxels. Pairwise Euclidian distances between all possible voxel pairs were calculated separately for ML, MSB and object-responsive regions in each hemisphere, then pooled across these regions after z-normalization within each region and hemisphere. To measure cortical distance between voxel pairs, we used center-to-center geometrical distance. To quantify the slope of the spatial fall-off pattern, we fitted a rational function with three free parameters [a/(1 + bx) + c] to the raw data using lsqcurvefit function in MATLAB. A likelihood ratio F-test was conducted to test whether the slope parameter b differed significantly from 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link